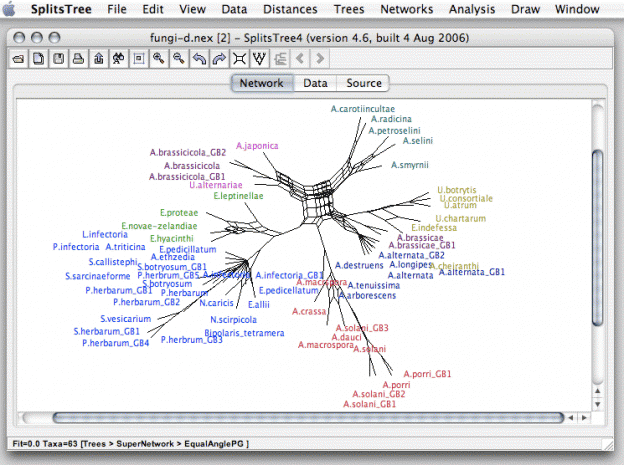
We’ve come a long way since proteins in pools by probability. Cliques provoked research into how best to validate their worth, which led to hamming distances, densitrees, splitstrees, and more.
My latest development milestone is the first working version of the perl code I’ve written for annotating proteins with their functions. I had a version of this code previously that did so for all cliques, but this time, I wanted to only handle the cliques indicated by Jim’s MatLab code.
The script works great, and outputs all cliques by binary signature, listing proteins and functions, and outputs the list of strains. Both on Octopussy and on my netbook, execution took just over an hour.
My next undertaking will be speeding the code up as much as possible, so that when I rework it to sit on the edwards.sdsu server with form submission, it will actually be usable in speed. The move to a web-based tool will of course include some other major headaches, but task one is getting the complexity analysis done and making strides towards speeding it up.