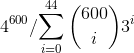How many possible bacterial species could there be with the usual species definition, and how many are there?
The paper “Scaling laws predict global microbial diversity” by Locey and Lennon suggests there are 1012 bacterial species based on the log normal distribution and ecological scaling laws. Does that make sense? How many possible bacterial species could there be on the planet?
Note: Bruno pointed out that it may be millions, not trillions:
@linsalrob After All, Only Millions?https://t.co/EClKiwBXNv
— Genomica Microbiana (@GenomicaMicrob) February 1, 2017
Check out these two papers:
We consider bacteria to be different species if their 16S genes are 97% different. However, the 16S genes are very highly conserved (which is why we can use them for metabarcoding). Here is the original paper describing the nine hypervariable regions in the 16S gene. As described in this paper, the nine regions span base pairs 69-99, 137-242, 433-497, 576-682, 822-879, 986-1043, 1117-1173, 1243-1294 and 1435-1465 for V1 through V9 using the E. coli 16S rRNA.
The E. coli 16S gene is 1543 bp long. The sum of the hypervariable regions is 565 bases.
Lets start with a couple of assumptions. As a reminder, in the Locey and Lennon paper they extrapolate their data over 30 orders of magnitude (that is from 100 to 1030). Therefore, we are not so concerned about exact numbers, and can round to easier numbers to help the math out. If you don’t believe me, watch this set of videos from Prof. Matt Anderson who shows why we should do this. (He’s a Physicist and should know!)
We assume that the 16S gene is 1500 bp long and has 900 bp of exactly identical sequences (the conserved region), and 600bp of different sequences (the variable regions). You can alter the numbers below to ignore the conserved regions if you don’t agree with my assumptions.
We need to know how many ways we can get 97% difference across that 600bp region. 97% difference means we need (1500bp * 0.03%) = 45 bp difference across the whole 16S gene, but of course it is concentrated in the 600 bp of difference.
The denominator is supposed to count the following: Given a specific sequence, how many distinct sequences differ from the given sequence by fewer than 45 sites on the 600 bp variable genome. For that reason the sum should technically run from 0 to 44. Also, once a mutational site is fixed, there are only 3 possible distinct mutations that differ from the given sequence. If two sites are fixed, there are nine, three sites fixed there are 27, etc. Therefore the factor should be 3i. So the full formula is:
4^{600}/{$$\sum_{i=0}^{44} {600 \choose i} 3^i}
The numerator is approximately 10400.
Although the denominator considers every option from 0 through 44, it is dominated by the 44: 600 choose 44 is 1067 and 344 is 1021. Therefore the denominator is about 1088.
Therefore, there are about 10312 possible bacterial species if every possible 16S DNA sequences was permuted and only those at 97% identity over 1,500 bases were considered distinct species.
Note that the EMP typically uses the 515f-806rB primer pair (e.g. here is Illumina’s protocol), and so this product is 292 bp long. We can do the same calculation with that product, assuming species have to be 97% different, we need 9 changes, therefore our equation becomes:
This becomes 10175 / (1015 * 104) = 10156.
If there really are only 1012 bacterial species, there is still a lot of 16S diversity out there that has not been sampled by biology!
By the way, Jim also put this into perspective: there have only been 1017 seconds since the big bang or the evolution of life (the big bang was 13.8 billion years ago and life evolved on earth ~3.6 billion (3.6 x 109 years ago; there are 2 x 107 seconds in a year). That means a new species evolves about every 105 seconds, which is basically one species per day!
Jim also pointed out that even though the denominator is an exact count (as the problem is defined), the quotient gives only a rough estimate of the number of distinguishable potential species. For example, all points within the unit sphere are closer than 1 unit from the center, but there are lots of pairs of points that are farther than 1 unit apart.
Edit!
This section below was the original solution, but Jim Nulton demonstrated that it is not correct! The correct solution is above, this is here for contemplation.
So our possible solutions are
4^{600}/{$$\sum_{i=1}^{45} {600 \choose i} 4^i}
The numerator is approximately 10400.
Although the denominator considers every option from 1 through 45, it is dominated by the 45: 600 choose 45 is 1068 and 445. is 1027. Therefore the denominator is about 10100.
Therefore, there are about 10300 possible bacterial species if every possible 16S DNA sequences was permuted and only those at 97% identity over 1,500 bases were considered distinct species.
Note that the EMP typically uses the 515f-806rB primer pair (e.g. here is Illumina’s protocol), and so this product is 292 bp long. We can do the same calculation with that product, assuming species have to be 97% different, we need 9 changes, therefore our equation becomes:
This becomes 10175 / (1016 * 105) = 10150.
If there really are only 1012 bacterial species, there is still a lot of 16S diversity out there that has not been sampled by biology!