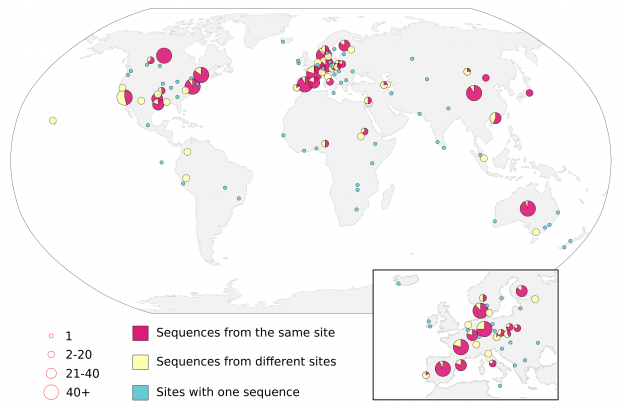
Making maps is hard. Even though we’ve been making maps for hundreds of years, it is still hard. Making good looking maps is really hard. We published a map that is both beautiful and tells a story, and this is the story of how we made that map.
But a figure like this does not appear immediately, it takes work to get something to look this good, and needless to say it wasn’t me that made it look so great!
Continue reading