So far I’ve learned some interesting things about services in Android. I also discovered that the issues with my build path (re: file manager troubles) are, well, known issues. Apparently the Android/Eclipse environment used to handle imports just fine, but nowadays it messes things up a bit when it builds .apk packages for phone deployment. A search on the Android Developers google group for ‘verify error’ turned up a detailed, though difficult for me to understand, explanation of exactly why Eclipse doesn’t build in method dependencies into apks. Either way, I’ve resolved myself to not importing methods – I’ll just cut and paste!
The AIDL tool for creating service interfaces is pretty cool, though a bit confusing to learn. From my current (limited) understanding, you can write Services which use Java primitives (and String… is that a primitive now?!), as well as any classes you write (provided you implement Parcelable!). As far as I’ve been able to discern, already-built classes like Hashtable are not in that list of usable classes. (Unless I change the class to implement parcelable, which I don’t have any real interest in doing…) Look for more on this tool when I have a better grasp of it – perhaps a future N&B article!
In my case, this unfortunately reduces me to the awkward approach of background threading for doing my JSON -> Hashtable conversion, list creation, and list sorting. Meanwhile, all http/cgi queries/uploads/downloads will be done in a background service. This approach isn’t terrible, however, as the difficult part to resume and recover would have been the download. If the background thread is interrupted, I can easily save the data and restart it when the app hits onResume(); Generally the time window on interrupting the sorting/list creation is a few seconds at most, so this will rarely come up.
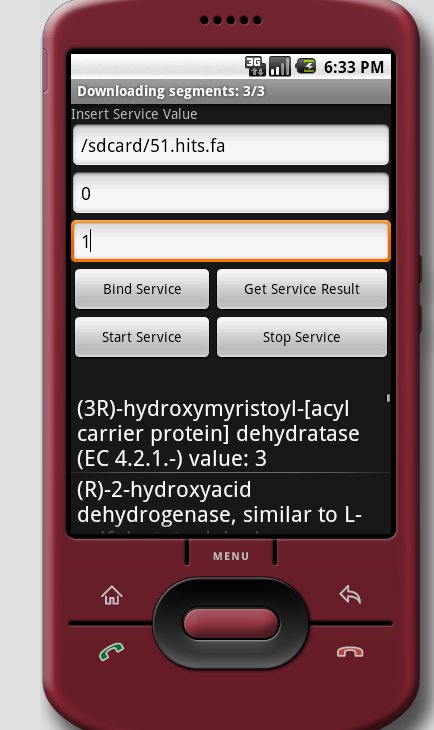
The other issue I’ve been struggling with today is that for some reason my download buffer is coming up as expecting 0 bytes to download when I run it from the service. Ugh, and that took me 1+ hours to track down even with the awesome Eclipse debugger. I had sort of hacked it into the service when I finally got my initial test running, so I went back and reconstructed everything from the pure MM classes (which still work). I now have a sort of pseudo-MM set up inside of my service test app, which I’ll get to test first thing tomorrow!
Edit, 8/18/09, Morning: It helps when you give your program good values to run tests on… (peek behind the read more)
Continue reading →